This guide explains how to migrate your tables from PostgreSQL/TimescaleDB into CrateDB. Without further ado, let's get into it.
Prerequisites
1. This tutorial uses PostgreSQL 13.2 with TimescaleDB 2.1.1 installed.
2. For the purpose of using a publicly available dataset in this tutorial, we'll be downloading one of the very useful sample datasets offered by Timescale: the so-called Devices Ops, which simulate readings from mobile devices. I'll be using the "devices_small" dataset.
3. If you want to also import the same dataset to PostgreSQL/TimescaleDB so you can follow this tutorial step by step, you can follow the instructions detailed in the Timescale documentation. (I actually imported the tables through pgAdmin though; if you want more details into how I did this, see [1] at the end).
Exporting the data from PostgreSQL
Note: in this section, we'll show you how to export your table as a CSV file. However, if you have more complex table structures (such as JSONB or array columns) you might prefer to export your data as JSON. If this is your case, see [2].
For this example, let's suppose we want to export the table "readings", included as a part of the "devices_small" dataset offered by Timescale. This is how it looks like:
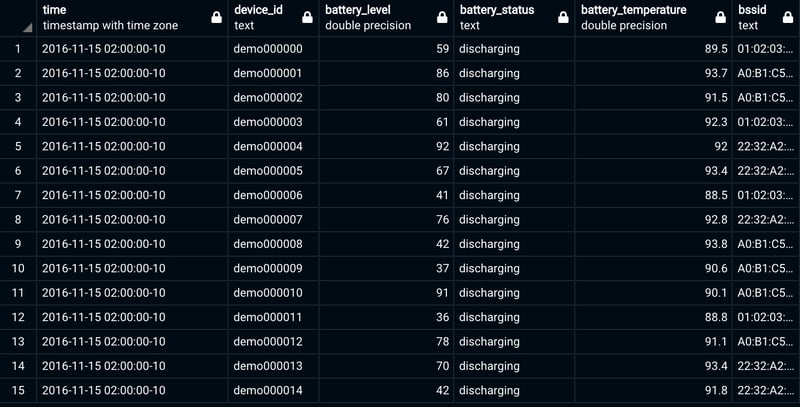
This table gives us the opportunity to make a first important point: if your dataset contains timestamps, you must transform these columns to the UNIX format in order to successfully import them to CrateDB.
In this case, we can do this by using EXTRACT FROM. We have two options:
- Option 1. To create a new table with the column corrected, exporting that table instead
In PostgreSQL, you can define a new table with the right format by running the following query:
CREATE TABLE readings_corrected AS
SELECT EXTRACT (epoch FROM time) * 1000 AS time_corrected, device_id, battery_level, battery_status, battery_temperature, bssid, cpu_avg_1min, cpu_avg_5min, cpu_avg_15min, mem_free, mem_used, rssi, ssid
FROM readings;
The result show like this:
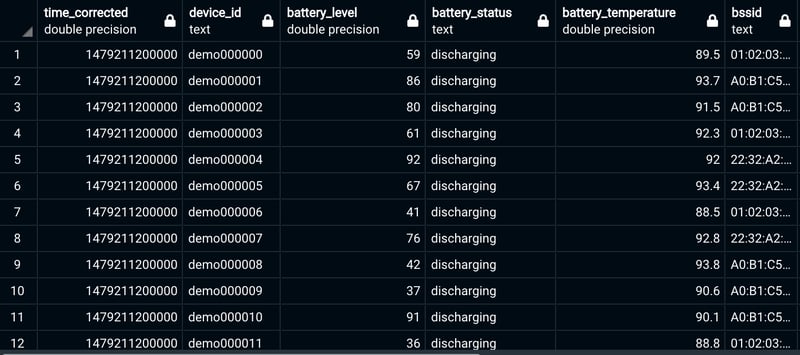
As you can see, the column "time_corrected" now comes with the format we were looking for.
Now, let's export this new table, which I have called "readings_corrected".
Using your terminal, use this command to connect to PostgreSQL, if you haven't done it yet:
psql -U postgres -d devices -h localhost
The corresponding syntax is psql -U <PostgreSQL user> -d <database> -h <host>. If needed, modify it accordingly to fit your user, database name, and so forth.
Next, run the code below to copy the table to a file in your Home folder. Again, modify it according to your needs, altering your table name, file name, and settings.
\copy (SELECT * FROM readings_corrected) TO '~/readings_corrected.csv' WITH (FORMAT 'csv', DELIMITER ',', QUOTE '"', ENCODING 'UTF-8', HEADER true)
- Option 2. To copy the columns directly from the command line
If you understand the previous process, you can skip the step of creating a new table in PostgreSQL by manually defining it column by column from the \copy command, also using EXTRACT FROM.
To export your table like that, follow these steps:
# Connect to PostgreSQL if you haven't done it yetpsql -U postgres -d devices -h localhost
# Define your table by naming the columns manually\copy (SELECT EXTRACT(epoch FROM "time") * 1000 AS "time", device_id, 'battery_level', battery_level, 'battery_status', battery_status, 'battery_temperature', battery_temperature, 'bssid', bssid, 'cpu_avg_1min', cpu_avg_1min, 'cpu_avg_15min', cpu_avg_15min, 'mem_free', mem_free, 'mem_used', mem_used, 'rssi', rssi FROM readings) TO '~/readings_corrected.csv' WITH (FORMAT 'csv', DELIMITER ',', QUOTE '"', ENCODING 'UTF-8', HEADER true)
Importing the data into CrateDB
Now, let's pretend we want to import the "readings_corrected" table to CrateDB.
First, define the table in the CrateDB Admin UI:
CREATE TABLE "readings_corrected"( time_corrected TIMESTAMP, device_id TEXT, battery_level DOUBLE PRECISION, battery_status TEXT, battery_temperature DOUBLE PRECISION, bssid TEXT, cpu_avg_1min DOUBLE PRECISION, cpu_avg_5min DOUBLE PRECISION, cpu_avg_15min DOUBLE PRECISION, mem_free DOUBLE PRECISION, mem_used DOUBLE PRECISION, rssi DOUBLE PRECISION, ssid TEXT)
You will see the table in CrateDB.
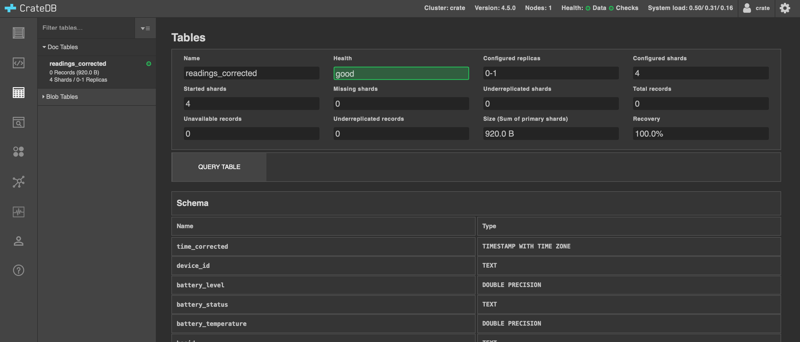
Now, we'll import the data using the COPY FROM command. In order for it to work, there are a few things to consider:
- The file with your data must be available to CrateDB. This means that if you want to import a file from your local computer, you first need to transfer the file to one of the CrateDB nodes.
- If you're running CrateDB inside a container, the file must also be inside it. For example, if you're using Docker, you may have to configure a Docker volume in order to do this; see [3] for a little bit more detail.
- In your COPY FROM statement, make sure you're using the appropriate URI scheme. In this tutorial, I’m using
file, but if your PostgreSQL instance is hosted on AWS RDS, you might want to consider the option of exporting directly to S3.
In my case, the COPY FROM statement had this shape:
COPY readings_corrected FROM 'file:///var/readings_corrected.csv'RETURN SUMMARY;
That's it! If there are no errors, the data is now in CrateDB:
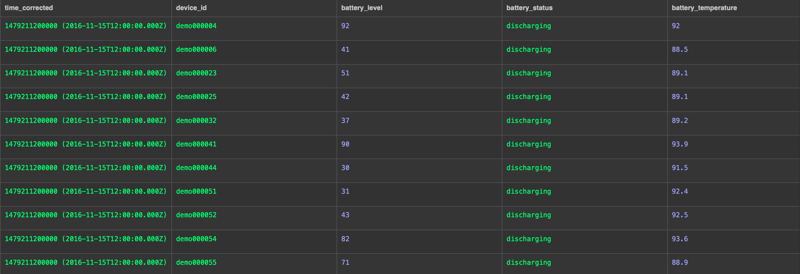
Additional notes
[1] How to import the "devices_small" dataset into PostgreSQL/TimescaleDB with pgAdmin
First, we have to create a database to host the data. Follow these steps:
# Connect to PostgreSQL using the superuser 'postgres'psql -U postgres -h localhost
# Create the database in PostgreSQL called 'devices'CREATE database devices;
# Add TimescaleDB to the databaseCREATE EXTENSION IF NOT EXISTS timescaledb;
Next, download the tar with the data. It includes two csv files and a sql file.
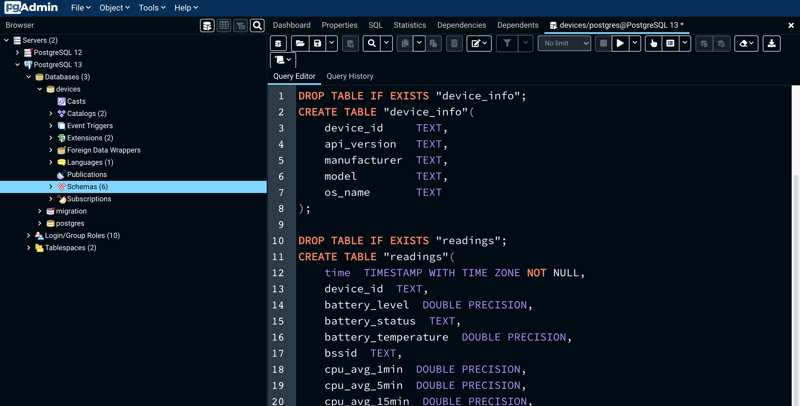
Before importing the csv files, first we need to define two tables in PostgreSQL/TimescaleDB: a "normal" table including some metadata ("device_info") and a hypertable with the mobile metrics ("readings"). In order to do so, run this (this is the statement provided by Timescale):
CREATE TABLE "device_info"( device_id TEXT, api_version TEXT, manufacturer TEXT, model TEXT, os_name TEXT);
CREATE TABLE "readings"( time TIMESTAMP WITH TIME ZONE NOT NULL, device_id TEXT, battery_level DOUBLE PRECISION, battery_status TEXT, battery_temperature DOUBLE PRECISION, bssid TEXT, cpu_avg_1min DOUBLE PRECISION, cpu_avg_5min DOUBLE PRECISION, cpu_avg_15min DOUBLE PRECISION, mem_free DOUBLE PRECISION, mem_used DOUBLE PRECISION, rssi DOUBLE PRECISION, ssid TEXT);
CREATE INDEX ON "readings"(time DESC);CREATE INDEX ON "readings"(device_id, time DESC);-- 86400000000 is in usecs and is equal to 1 daySELECT create_hypertable('readings', 'time', chunk_time_interval => 86400000000);
Once the tables are created, right-click on them, selecting the "Import/Export" function. A window like this will display:
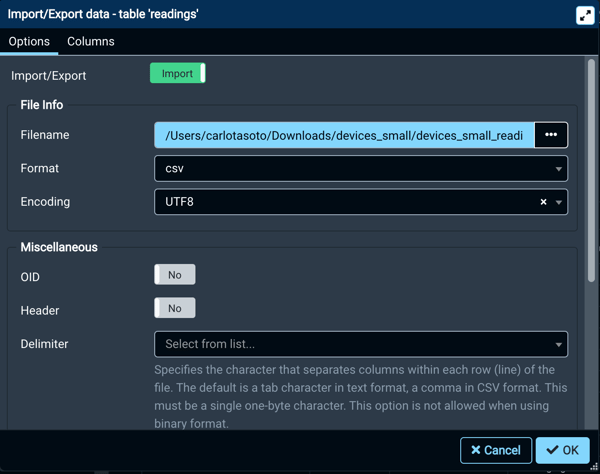
Once you're done, press "OK". You can check if everything has worked by taking a look at the tables.
[2] How to export the data from PostgreSQL as JSON
You can use:
# Connect to PostgreSQL if you haven't done it yetpsql -U postgres -h localhost
# Create the JSON object, transforming the time columns into UNIX format\copy (SELECT JSON_BUILD_OBJECT('time', EXTRACT(epoch FROM "time") * 1000, 'device_id', device_id, 'battery_level', battery_level, 'battery_status', battery_status, 'battery_temperature', battery_temperature, 'bssid', bssid, 'cpu_avg_1min', cpu_avg_1min, 'cpu_avg_15min', cpu_avg_15min, 'mem_free', mem_free, 'mem_used', mem_used, 'rssi', rssi ) FROM readings) TO '~/device_readings.json' WITH (FORMAT 'text')
The generated JSON will look like this:
{"time" : 1479211200000, "device_id" : "demo000000", "battery_level" : 59, "battery_status" : "discharging", "battery_temperature" : 89.5, "bssid" : "01:02:03:04:05:06", "cpu_avg_1min" : 24.81, "cpu_avg_15min" : 8.654, "mem_free" : 410011078, "mem_used" : 589988922, "rssi" : -50}
A few other notes on the topic of more complex table definitions:
About arrays/JSONB
Both arrays and JSONB columns can be included in the generated JSON document by specifying a corresponding key and the column name.
For example, consider this table:
CREATE TABLE readings ("time" TIMESTAMP, memory FLOAT[], cpu JSONB);INSERT INTO readings VALUES ('2016-11-15T12:00:00+00:00', '{650609585, 349390415}', '{"cpu_avg_1min": 5.26,"cpu_avg_5min": 6.172, "cpu_avg_15min": 6.51066666666667}');
To export this as JSON, use:
# Connect to PostgreSQL if you haven't done it yetpsql -U postgres -h localhost
# Use JSON_BUILD_OBJECT to build the documentSELECT JSON_BUILD_OBJECT('time', EXTRACT(epoch FROM "time") * 1000, 'memory', memory, 'cpu', cpu) FROM readings;
The output will look as follows:
{"time" : 1479211200000, "memory" : [650609585,349390415], "cpu" : {"cpu_avg_1min": 5.26, "cpu_avg_5min": 6.172, "cpu_avg_15min": 6.51066666666667}}
And in CrateDB, the target table would be defined like this:
CREATE TABLE readings ( "time" TIMESTAMP, memory INTEGER[], cpu OBJECT AS ( cpu_avg_1min FLOAT, cpu_avg_5min FLOAT, cpu_avg_15min FLOAT ));
Geo points
For example, consider:
CREATE TABLE points (p point);INSERT INTO points VALUES ((1,2));
ROW_TO_JSON will serialize the record as {"p":"(1,2)"}, which cannot be parsed by CrateDB. Instead, we can use JSON_BUILD_ARRAY to transform the point into an array:
SELECT JSON_BUILD_OBJECT('p', JSON_BUILD_ARRAY(p[0], p[1])) FROM geo i;
The generated JSON will now be understood by CrateDB for its proper importing.
[3] How to make files available to a Docker container
Let's suppose you use the following command to access CrateDB through Docker:
docker run --rm --name=cratedb --publish=4200:4200 --publish=5432:5432 --volume="$(pwd)/var/lib/crate:/data" crate
You can make the file available to the container by running a command similar to the one below:
docker cp /Users/carlotasoto/readings_corrected.csv a4298201d59:var
This is the syntax:
docker cp /hostfile (container_id):/(to_the_place_you_want_the_file_to_be))
In my case, “a4298201d59” was the container id. You can reveal your container id with docker ps.